van Hazel I, Santini F, Müller J, Chang BS
BMC Evol. Biol. 2006;6:97
PubMed PMID: 17107620
Abstract
BACKGROUND: Vertebrate SWS1 visual pigments mediate visual transduction in response to light at short wavelengths. Due to their importance in vision, SWS1 genes have been isolated from a surprisingly wide range of vertebrates, including lampreys, teleosts, amphibians, reptiles, birds, and mammals. The SWS1 genes exhibit many of the characteristics of genes typically targeted for phylogenetic analyses. This study investigates both the utility of SWS1 as a marker for inferring vertebrate phylogenetic relationships, and the characteristics of the gene that contribute to its phylogenetic utility.
RESULTS: Phylogenetic analyses of vertebrate SWS1 genes produced topologies that were remarkably congruent with generally accepted hypotheses of vertebrate evolution at both higher and lower taxonomic levels. The few exceptions were generally associated with areas of poor taxonomic sampling, or relationships that have been difficult to resolve using other molecular markers. The SWS1 data set was characterized by a substantial amount of among-site rate variation, and a relatively unskewed substitution rate matrix, even when the data were partitioned into different codon sites and individual taxonomic groups. Although there were nucleotide biases in some groups at third positions, these biases were not convergent across different taxonomic groups.
CONCLUSION: Our results suggest that SWS1 may be a good marker for vertebrate phylogenetics due to the variable yet consistent patterns of sequence evolution exhibited across fairly wide taxonomic groups. This may result from constraints imposed by the functional role of SWS1 pigments in visual transduction.

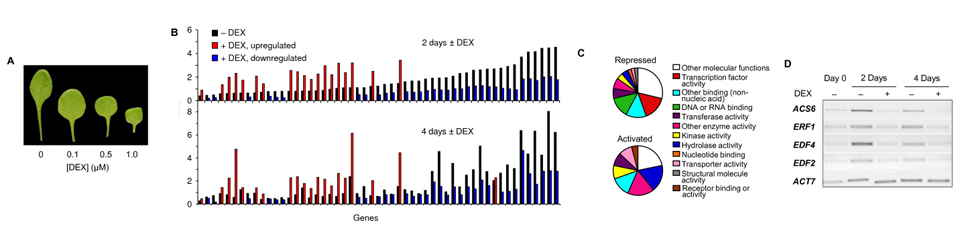 Lumba S, Tsuchiya Y, Delmas F, Hezky J, Provart NJ, Shi Lu Q, McCourt P, Gazzarrini S
Lumba S, Tsuchiya Y, Delmas F, Hezky J, Provart NJ, Shi Lu Q, McCourt P, Gazzarrini S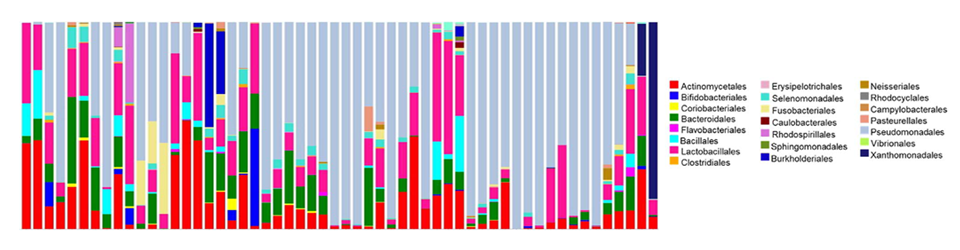 Maughan H, Cunningham KS, Wang PW, Zhang Y, Cypel M, Chaparro C, Tullis DE, Waddell TK, Keshavjee S, Liu M, Guttman DS, Hwang DM
Maughan H, Cunningham KS, Wang PW, Zhang Y, Cypel M, Chaparro C, Tullis DE, Waddell TK, Keshavjee S, Liu M, Guttman DS, Hwang DM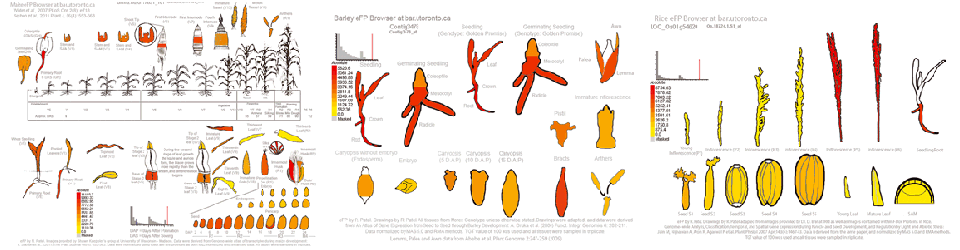 Patel RV, Nahal HK, Breit R, Provart NJ
Patel RV, Nahal HK, Breit R, Provart NJ